Genome-wide comparison and identification of myosin gene family in Arabidopsis thaliana and Helianthus annuus
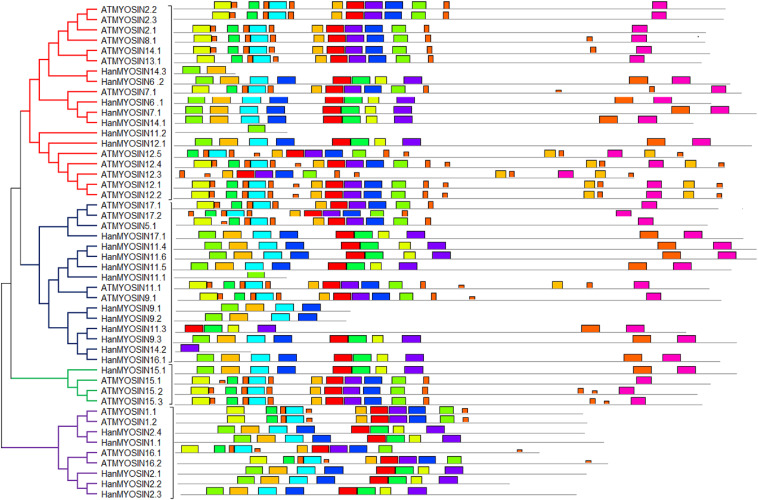
Myosins are essential components of organelle trafficking in all the eukaryotic cells. Myosin driven movement plays a vital role in the development of pollen tubes, root hairs and root tips of flowering plants. The present research characterized the myosin genes in Arabidopsis thaliana and Helianthus annuus by using different computational tools. We discovered a total of 50 myosin genes and their splice variants in both pant species. Phylogenetic analysis indicated that myosin genes were divided into four subclasses. Chromosomal location revealed that myosin genes were located on all five chromosomes in A. thaliana, whereas they were present on nine chromosomes in H. annuus. Conserved motifs showed that conserved regions were closely similar within subgroups. Gene structure analysis showed that Atmyosin2.2 and Atmyosin2.3 had the highest number of introns/exons. Gene ontology analysis indicated that myosin genes were involved in vesicle transport along actin filament and cytoskeleton trafficking. Expression analysis showed that expression of myosin genes was higher during the flowering stage as compared to the seedling and budding stages. Tissue specific expression indicated that HanMYOSIN11.2, HanMYOSIN16.2 were highly expressed in stamen, whereas HanMYOSIN 2.2, HanMYOSIN 12.1 and HanMYOSIN 17.1 showed higher expression in nectary. This study enhance our understanding the function of myosins in plant development, and forms the basis for future research about the comparative genomics of plant myosin in other crop plants.