A collaborative resource to build consensus for automated left ventricular segmentation of cardiac MR images
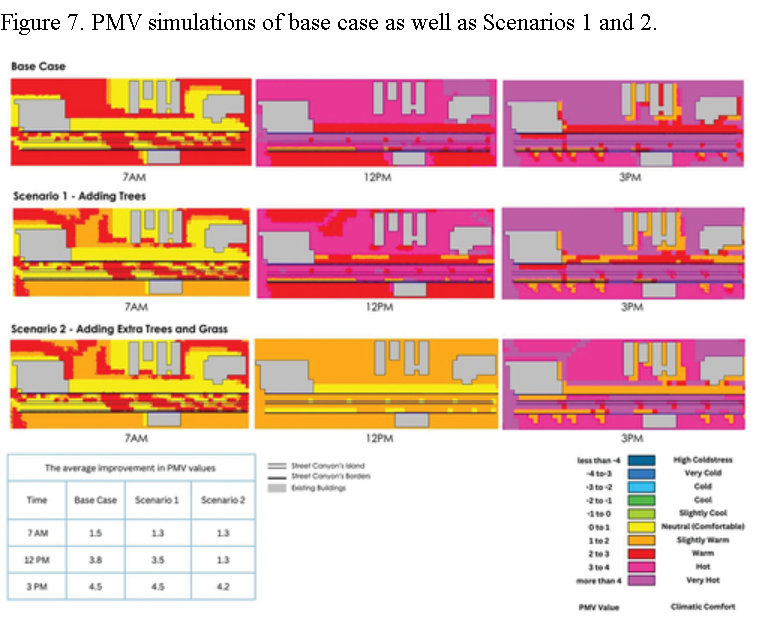
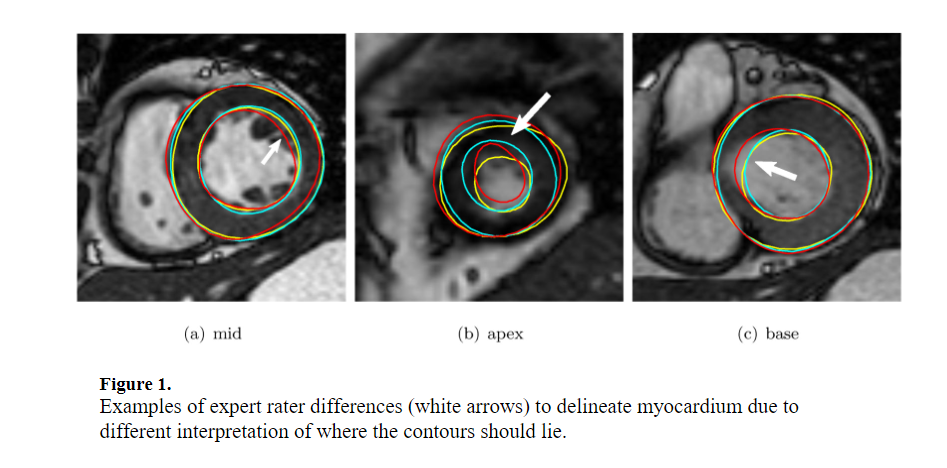
A collaborative framework was initiated to establish a community resource of ground truth segmentations from cardiac MRI. Multi-site, multi-vendor cardiac MRI datasets comprising 95 patients (73 men, 22 women; mean age 62.73. ±. 11.24. years) with coronary artery disease and prior myocardial infarction, were randomly selected from data made available by the Cardiac Atlas Project ( Fonseca et al., 2011). Three semi- and two fully-automated raters segmented the left ventricular myocardium from short-axis cardiac MR images as part of a challenge introduced at the STACOM 2011 MICCAI workshop ( Suinesiaputra et al., 2012). Consensus myocardium images were generated based on the Expectation-Maximization principle implemented by the STAPLE algorithm ( Warfield et al., 2004). The mean sensitivity, specificity, positive predictive and negative predictive values ranged between 0.63 and 0.85, 0.60 and 0.98, 0.56 and 0.94, and 0.83 and 0.92, respectively, against the STAPLE consensus. Spatial and temporal agreement varied in different amounts for each rater. STAPLE produced high quality consensus images if the region of interest was limited to the area of discrepancy between raters. To maintain the quality of the consensus, an objective measure based on the candidate automated rater performance distribution is proposed. The consensus segmentation based on a combination of manual and automated raters were more consistent than any particular rater, even those with manual input. The consensus is expected to improve with the addition of new automated contributions. This resource is open for future contributions, and is available as a test bed for the evaluation of new segmentation algorithms, through the Cardiac Atlas Project ( www.cardiacatlas.org). © 2013 Elsevier B.V.